Bioprocessing References
- Baeshen NA, Baeshen MN, Sheikh A, et al. Cell factories for insulin production. Microb Cell Fact. 2014;13:141. doi: 10.1186/s12934-014-0141-0
- Bell et al. 2021, Characterization of a novel high-throughput, high-speed and high-precision plate-based image cytometric cell counting method, Cell and Gene Therapy Insight
- Dahodwala H, Lee KH. The fickle CHO: a review of the causes, implications, and potential alleviation of the CHO cell line instability problem. Curr Opin Biotech. 2019;60:128-37. doi: 10.1016/j.copbio.2019.01.011
- Dalton AC, Barton WA. Over-expression of secreted proteins from mammalian cell lines. Protein Sci. 2014;23:517-525. doi: 10.1002/pro.2439
- Fischer S, Handrick R, Otte K. The art of CHO cell engineering. A comprehensive retrospect and future perspectives. Biotechnol Adv. 2015;33:187801896. doi: 10.1016/j.biotechadv.2015.10.015
- Gupta SK, Shukla P. Sophisticated cloning, fermentation, and purification technologies for an enhanced therapeutic protein production: A Review. Front Pharmacol. 2017;8:419. doi: 10.3389/fphar.2017.00419
- Hansen HG, Nilsson CN, Lund AM, et al. Versatile microscale screening platform for improving recombinant protein productivity in Chinese hamster ovary cells. Sci Rep. 2015;5:18016. doi: 10.1038/srep18016
- International Organization for Standardization. Biotechnology – Cell counting – Part 1: General guidance on cell counting methods. (Standard No. 20391-1:2018). Available at: https://www.iso.org/standard/68879.html
- International Organization for Standardization. Biotechnology – Cell counting – Part 2: Experimental design and statistical analysis to quantify counting method performance. (Standard No. 20391-2:2019). Published 2019. Available at: https://www.iso.org/standard/67892.html
- Joubert S, Dodelet V, Béliard R, Durocher Y. Biomanufacturing of monoclonal antibodies. Med Sci (Paris). 2019;35(12):1153-1159. doi: 10.1051/medsci/2019219
- Lee JS, Kallehauge TB, Pedersen LE, Kildegaard HF. Site-specific integration in CHO cells mediated by CRISPR/Cas9 and homology-directed DNA repair pathway. Sci Rep. 2015;5:8572. doi: 10.1038/srep08572
- Lund AM, Kildegaard HF, Petersen MB, et al. A versatile system for USER cloning-based assembly of expression vectors for mammalian cell engineering. PLoS One. 2014;9:e96693. doi: 10.1371/journal.pone.0096693
- Pierce et al. 2021, Outcomes from a cell viability workshop: fit-for-purpose considerations for cell viability measurements for cellular therapeutic products, Cell and Gene Therapy Insight
- Ritacco FV, Wu Y, Khetan A. Cell culture media for recombinant protein expression in Chinese hamster ovary (CHO) cells: History, key components, and optimization strategies. Biotechnol. Prog. 2018;34:1407-1426. doi: 10.1002/btpr.2706
- Ronda C, Pedersen LE, Hansen HG, et al. Accelerating genome editing in CHO cells using CRISPR Cas9 and CRISPy, a web-based target finding tool. Biotech Bioeng. 2014;111:1604-1616. doi: 10.1002/bit.25233
- Shaw D, Yim M, Tsukuda J, et al. Development and characterization of an automated imaging workflow to generate clonally-derived cell lines for therapeutic proteins. Biotechnol Prog. 2018;34:584-592. doi: 10.1002/btpr.2561
- Tharmalingam T, Barkhordarian H, Tejeda N, et al. Characterization of phenotype and genotypic diversity in sublclones derived from a clonal cell line. Biotechnol Prog. 2018;34:613-623. doi: 10.1002/btpr.2666
- World Health Organization. Recommendations for the evaluation of animal cell cultures as substrates for the manufacture of biological medicinal products and for the characterization of cell banks. WHO Expert Committee on Biological Standardization: sixty-first report; Annex 3. WHO Technical Report Series No. 978, 2013. Available at https://www.who.int/biologicals/vaccines/TRS_978_Annex_3.pdf
- World Health Organization. Guidelines on evaluation of monoclonal antibodies as similar biotherapeutic products (SBPs); Annex 2. WHO Technical Report Series No. 1004, 2016. Available at https://www.who.int/biologicals/biotherapeutics/WHO_TRS_1004_web_Annex_2.pdf?ua=1
- Yamane-Ohnuki N, Kinoshita S, Inoue-Urakubo M, et al. Establishment of FUT8 knockout Chinese hamster ovary cells: An ideal host cell line for producing completely defucosylated antibodies with enhanced antibody-dependent cellular cytotoxicity. Biotechnol Bioeng. 2004;87:614-622. doi: 10.1002/bit.20151
- Zhang H, Chan LL, Rice W, et al. Novel high-throughput cell-based hybridoma screening methodology using the Celigo Image Cytometer. J Immunol Methods. 2017;447:23-30. doi: 10.1016/j.jim.2017.04.003
- Zhou Y, Shaw D, Lam C, et al. Beating the odds: The Poisson distribution of all input cells during limiting dilution grossly underestimates whether a cell line is clonally-derived or not. Biotechnol Prog. 2018;34:559-569. doi: 10.1002/btpr.2560
- Zigon ES, Purseglove SM, Toxavidis V, et al. A rapid single cell sorting verification method using plate-based image cytometry. Cytometry A. 2018;93:1060-1065. doi: 10.1002/cyto.a.23520
Quality control to develop high-quality cell cultures
- Rapid cell counting
- Accurate assessment of proliferation and viability
- Ensuring monoclonality
- Functional and screening assays
Important criteria for clone selection [Gupta and Shukla, 2017]
- Cell growth pattern
- High protein expression
- Appropriate PTMs
- Genetic stability
Bioprocessing References
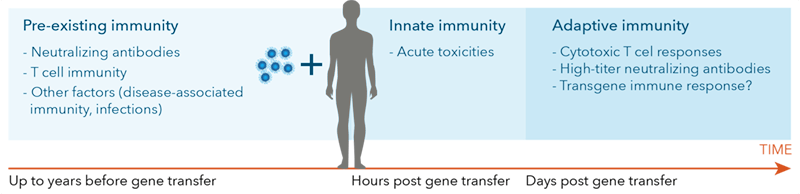
Figure 1. Wild-type AAV exposure and patient characteristics determine an individual’s response at vector delivery and overtime.
It is critical to assess immune status in patients to monitor gene therapy efficacy [33]. Subjects previously treated with gene therapy may require a second dose as an initial treatment may have activated a patient’s adaptive immune system to eliminate the vector upon reintroduction. This proves that assessing a patient’s immune status is critical to monitoring gene therapy efficacy [33].
To improve the effectiveness of vector delivery, image cytometers can be employed for:
- Rapid screening for nAbs prior to treatment
- Cell-based in vitro transduction inhibition assays
- Long-term patient monitoring
- Humoral immune responses for antibodies against the vector and gene therapy protein product
- T-cell immune responses
Use plate-based imaging to screen patient sera for neutralizing antibodies and monitoring immune responses over time
The Celigo Image Cytometer is a plate-based high-throughput system that simultaneously images and analyzes cells in standard multi-well plates using brightfield and fluorescence technology. The Celigo rapidly screens for sera nAbs that can neutralize AAV vectors for potential pre-existing immunities.
- High-throughput automated imaging and direct analysis in 96-well plates
- No trypsinization necessary to count cells
- Measure antibody neutralization effects in less than ten minutes
This section provides two technical examples of how the Celigo Image Cytometer can be used to develop a robust screening assay for potential pre-existing immunity to AAV vectors.