Normalization of Agilent Seahorse™ XF Data with Direct Cell Counting Using the Celigo Image Cytometer
In the recent years, the Seahorse™XF Analyzer has been diligently utilized to measure the Oxygen Consumption Rate (OCR) and Extracellular Acidification Rate (ECAR)/ Proton Production Rate (PPR) for understanding energy metabolism and cellular functions [1].
One of the important steps in performing Seahorse™ XF cell metabolism analysis is normalization of the resultant data to minimize inconsistency and variations from well-to-well due to potential non-uniform cell seeding across the microplate. In general, normalization has been commonly performed using total protein amount in each well such as BCA assays [2].
There are some known issues with protein-based normalization method
- Cells can be lost during the assay due to the metabolic inhibitors used during the assay
- Normalizing Seahorse™ data using total protein analysis is time-consuming
- It requires multiple steps such as cell lysing, incubation, and transferring lysed content for analysis
- Washing and transfer procedure can introduce risk such as cell loss or pipetting error, thus skewing the final normalized results
- The BCA assay can take up to ~45 min for the entire 96 well plate
An alternative method that can improve Seahorse™ XF normalization is performing direct cell count per well prior to the metabolic stress tests.
Direct cell count-based normalization method eliminates issues from protein analysis
- Cell counting is a non-destructive method to generate data for Seahorse™ normalization
- It can take account of non-uniform seeding density across the well and plate
- Downstream assays are still possible after normalization
- Direct cell counting can significantly reduce the amount of time required to perform normalization
Using Celigo Image Cytometer for whole well direct cell counting
In recent years, Celigo Image Cytometer has been used to perform bright field or fluorescence-based direct cell counting for Seahorse™ XF data normalization [3]. Due to its ability to capture whole-well images, at high speed; Celigo direct cell counting application has been adapted into the general workflow for Seahorse™ XF96, XFe96, and XF24 Analyzers.
Celigo Image Cytometer provides a fast, effective, and cell stress-free technique to count cells plated onto uncoated or coated culture plates. The system contains built-in plate profiles for the Seahorse™ plates and will acquire one whole well image of a 96-well plate in ~4-5 seconds, and analysis of the entire plate is ~6 min in bright field, and ~9 min in fluorescence.
Celigo Image Cytometer can be used to perform direct cell counting of suspension and adherent cells in bright field, as well as fluorescence such as Hoechst 33342 or Live-nuclear dyes staining. Both bright field and fluorescence methods can be used effectively to normalize Seahorse™ XF data.
The use of live nuclear dyes (Nexcelom Bioscience) can emit a brighter fluorescence signal, and have little to no cytotoxic & cytokinetic effects. In addition, the cell identification algorithms are highly accurate for segmenting suspension or adherent cells for both bright field and fluorescence imaging. Fluorescence live nuclear dyes can improve accuracy for counting adherent cells that tend to flatten out after attachment.
Furthermore, Celigo has a large dynamic range of counting cells in whole well, i.e. counting 1 to ~120,000 cells per well for suspension cells and 1 to ~90,000 cells for adherent cells in a 96-well plate.
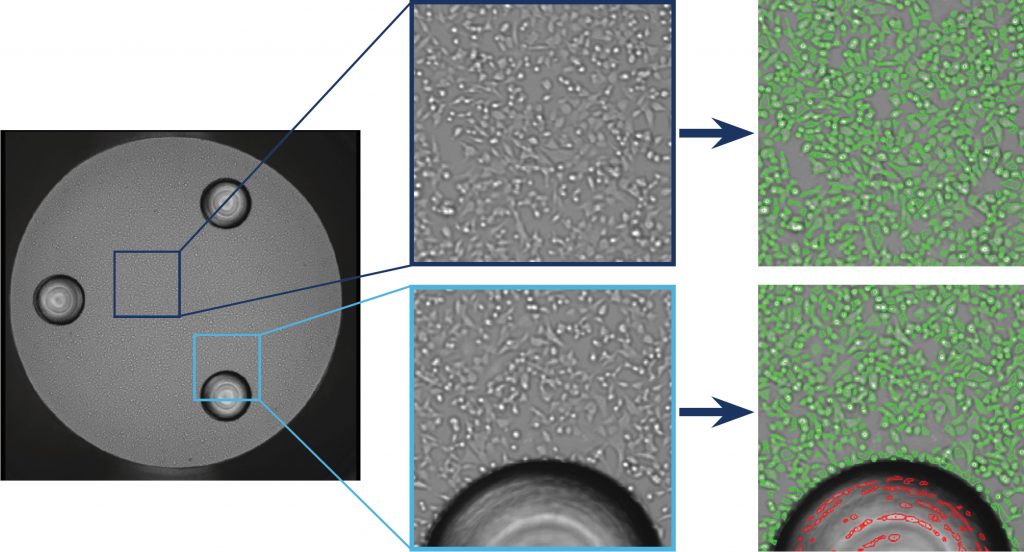

Figure. 1 Whole-well images captured by Celigo from a Seahorse™ XF well with 3 posts. Accurate direct cell count is shown where cells are circled in green and exclusion of the post is shown in red.
Bright field-based direct cell counting method for Seahorse™ XF normalization
In this experiment, the glycolytic function (ECAR) was assessed for DCIS and HCT116 cell lines for approximately 160 min. The Seahorse™ XF data were normalized to both protein amount and bright field cell counting using the Celigo Image Cytometer. The normalized time-dependent ECAR data were then compared, which showed improvement in cell line differentiation using the cell count normalization.
- DCIS and HCT116 cells were seeded at 12,500 cells/well and 10,000 cells/well, respectively, in the Seahorse™ XF 96-well plate and incubated overnight
- The plate was first visualized on microscope for approximately 95% confluence
- The confluence was then confirmed at 98% using the Celigo, as well as QC for cell health, morphology, seeding uniformity and purity (no contamination) prior to washing
- Next, washed with the appropriate assay media
- Subsequently, the plate was imaged and analyzed on the Celigo using direct cell counting application
- Next, the plate is placed in a 37° C incubator without CO2 for 45 minutes
- The plate was then analyzed using Seahorse XFe96 to measure ECAR for 160 min
- Finally, the plate was used to perform protein analysis for normalization
The Celigo captured bright field images are displayed in Figure 1, which showed a whole well image of a Seahorse™ XF well with the 3 posts. In addition, the zoomed in images showed accurate direct cell counting of the HC116 cells (green outline), as well as exclusion of the post area (red outline).
By utilizing the graphing function and well mask feature in the Celigo software, researchers can count cells at specific percentages of area within the well, which may allow better normalization for the Seahorse™ XF data (Figure 2).
The normalized ECAR time-dependent data for protein and cell count normalization are shown in Figure 3. It is obvious that measurements with protein normalization cannot distinguish between the glycolytic function of DCIS and HCT116 cell lines. In contrast, the cell count normalization method showed a reduction in standard deviation as well as a clear separation between the ECAR values of DCIS and HCT116 cell lines.
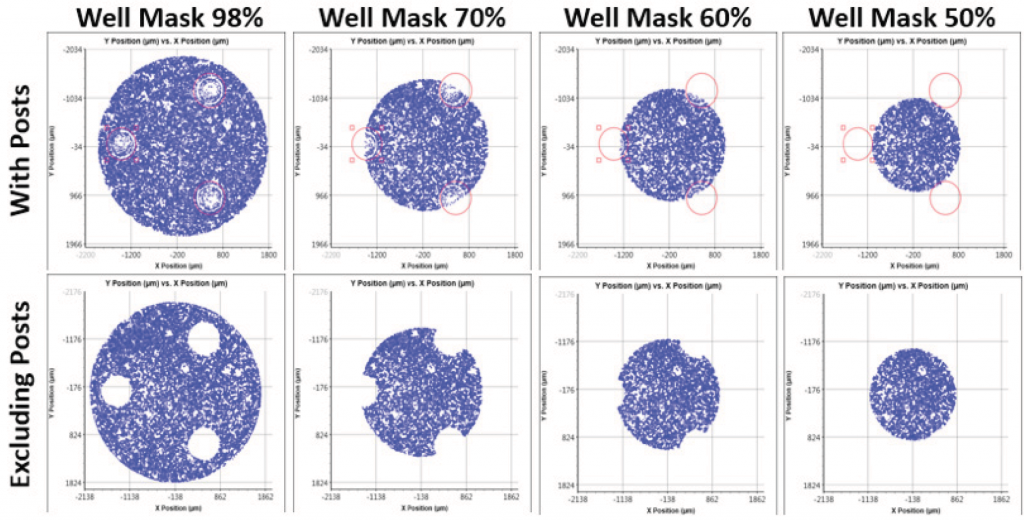
Figure. 2 The graphing and well mask function of the Celigo software allows for cell counts at specific percentages within the area of a well.
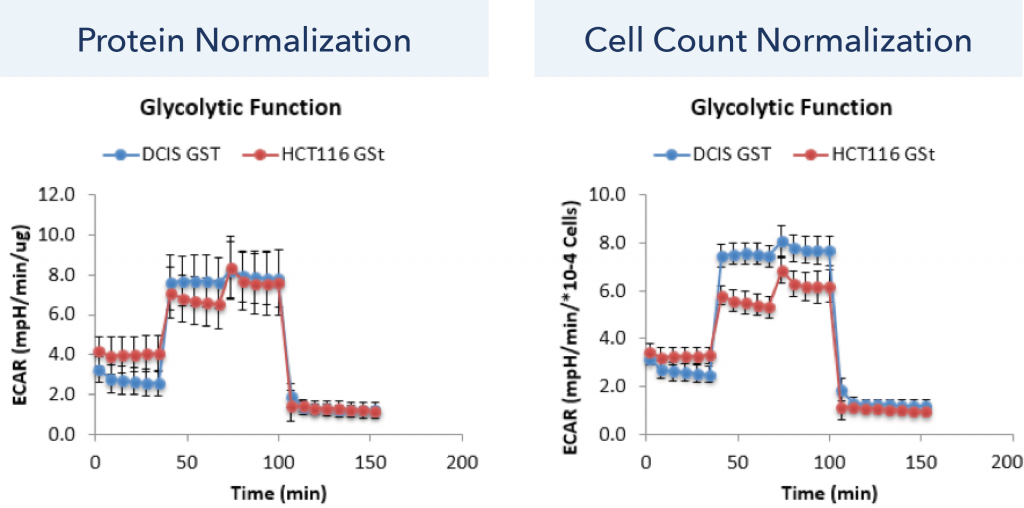
Figure. 3 Normalized ECAR time-dependent data for protein and cell count normalization.
Fluorescence-based direct cell counting method for Seahorse™ XF normalization
In this experiment, both mitochondrial respiration (OCR) and glycolytic function (ECAR) were assessed for T cells for approximately 60 min. The Seahorse™ XF data were normalized to both protein amount and fluorescent cell counting using the Celigo Image Cytometer. The normalized endpoint ECAR data were then compared, which showed improvement in drug response differentiation using the cell count normalization.
- T cells were seeded at 100,000 cells/well in the Seahorse™ XF 96-well plate
- The cells were allowed to settle to the bottom of the plate
- The plate was then placed into the Seahorse™ XF Analyzer and allowed delivery of Hoechst 33342, and subsequently analyzed for 60 min
- The plate was then counted on the Celigo for total Hoechst positive cells
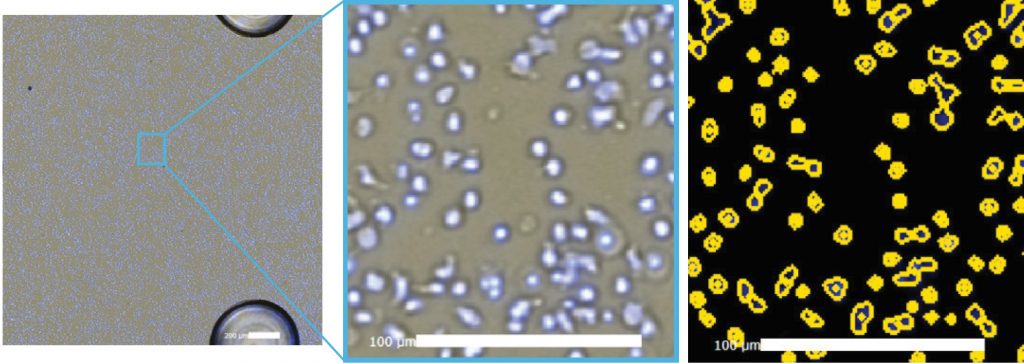
Figure. 4 Celigo captured bright field image (left), zoomed merged bright field and Hoechst image (center), and yellow ROI counted Hoechst stained cells (right).
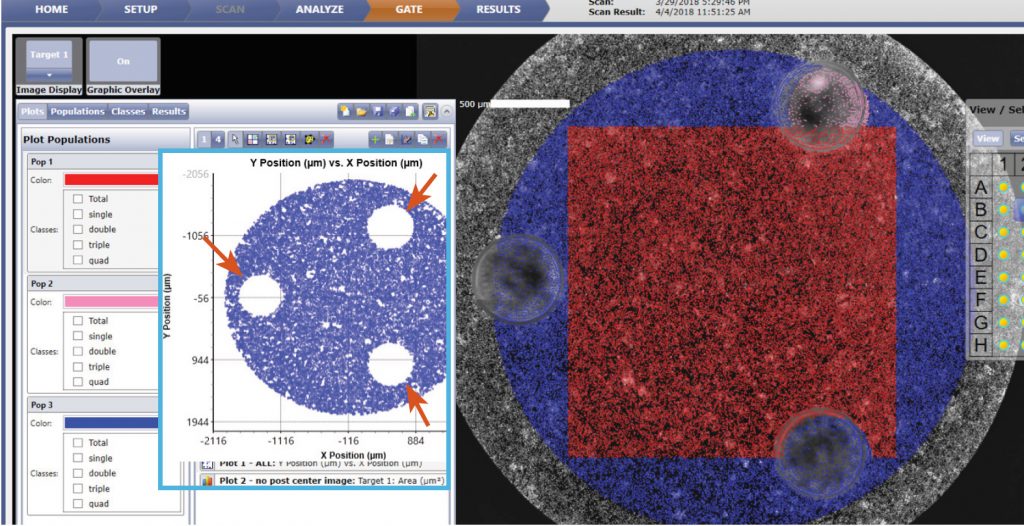
Figure. 5 Using the Celigo gating function, the location graph is generated to gate out the 3 posts (indicated by red arrows above) in the well using fluorescence. In addition, if only the area between the 3 posts are analyzed (shown with red square), scanning and counting require only 2 min per plate.
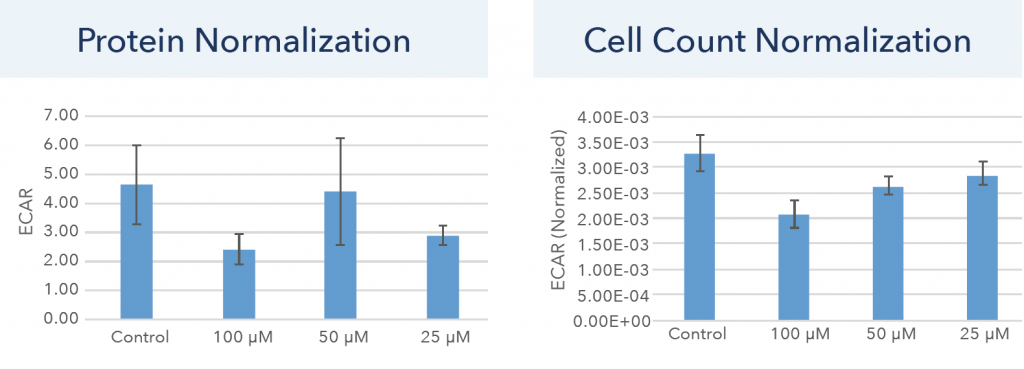
Figure. 6 Normalized ECAR endpoint data for protein and cell count normalization.
The Celigo captured bright field and Hoechst fluorescence overlay images are shown in Figure 4. The counted image showed yellow outlines indicating counted Hoechst positive cells. Similarly, under the gating function, the location graph can be generated to gate out the 3 posts in the well using fluorescence (Figure 5).
The normalized ECAR endpoint data for protein and cell count normalization are shown in Figure 6. Similarly, the protein and cell count normalization method showed distinct results. Using protein normalization, large standard deviation for each sample can be observed, as well as inexplicable drug treatment trends. On the other hand, the cell count normalization showed a reduction in standard deviation, as well as showing expected drug treatment results with a decrease in ECAR as drug concentration increased.
Speed to Results
The Celigo Image Cytometer can scan whole-well images and directly count cells in less than 6 and 9 min per 96-well plate (less than 4 and 5 seconds per well) using bright field and fluorescence imaging, respectively (Figure 7). If only the area between the 3 posts are captured, the entire process requires only 2 min. Researchers investigating mitochondrial respiration and glycolytic functions using Seahorse™ XF Analyzer can rapidly and effectively normalize the OCR and ECAR measurements without additional procedures like protein analysis.
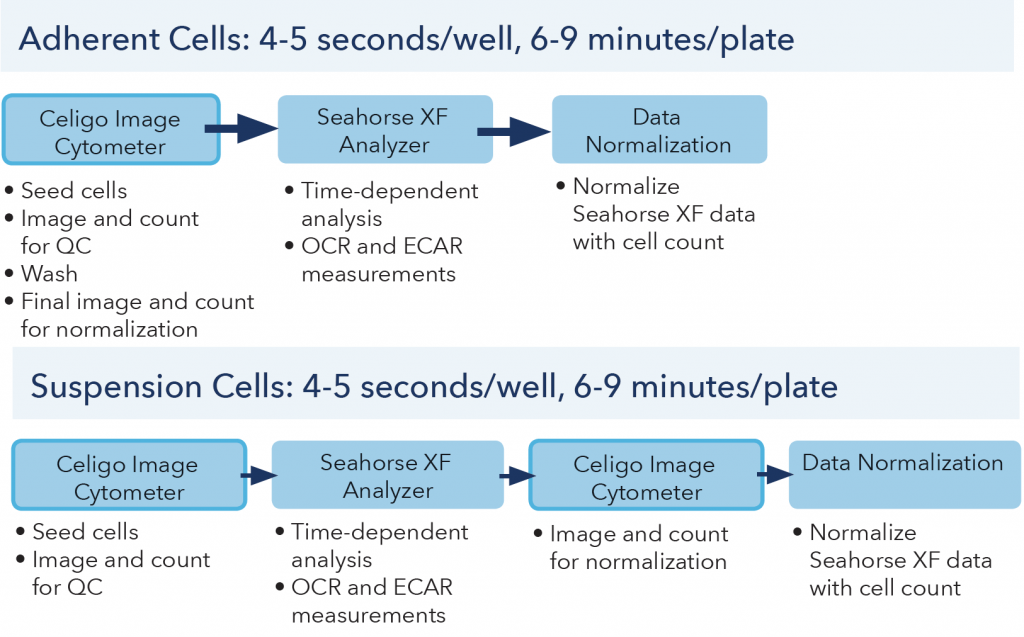
Figure. 7 Celigo workflows for adherent and suspension cells.
Whole Well Image and Image Quality
Image quality is a critical advantage for the Celigo Image Cytometer. Because of components of F-Theta lens and Galvanometric Mirror, Celigo generates highly uniform and rapid whole well imaging in both bright field and fluorescence for the entire plate. Therefore, it is unnecessary to extrapolate from one single image in the middle of the well, which may create uncertainty and inaccurate counting in the well if the cells are not uniformly seeded (Figure 8). These images can also help to identify issues with cells or incorrect seeding if assay results are not as expected.
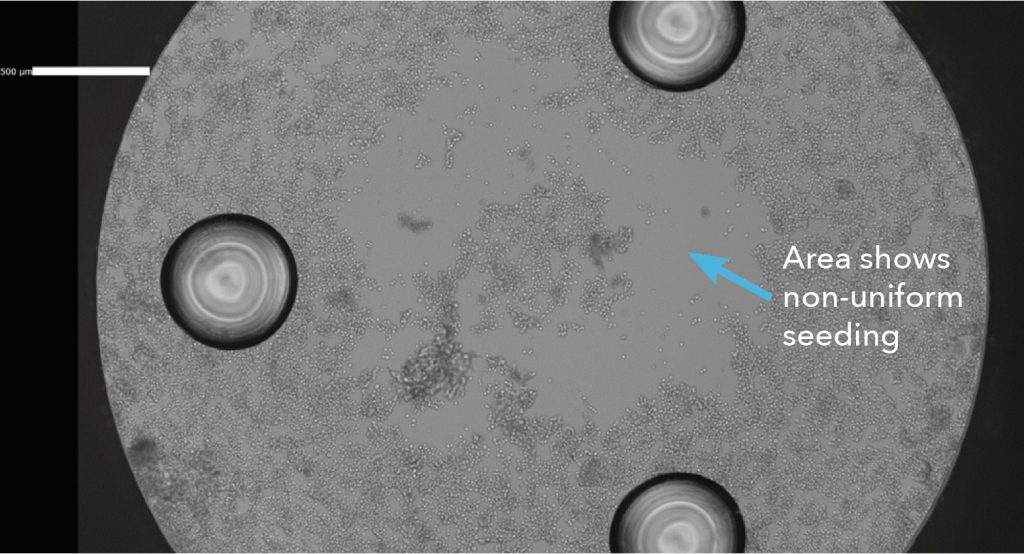

Figure. 8 Celigo whole-well bright field image shows non-uniformly seeded cells.
Sophisticated and Accurate Cell Counting Algorithm
Since cells come in all shapes and sizes, researchers also require a flexible and sophisticated cell counting platform to ensure accurate results for normalization. Both adherent and suspension cells can utilize a variety but specific counting algorithm to count the target cells in both bright field and fluorescent images. In addition, location plots are used to specifically gate out the 3 post areas to further improve cell counting accuracy (Figure 2).
Perform label-free direct cell counting
The Celigo Image Cytometer can be used to directly count both adherent and suspension cells without fluorescent labels, which can eliminate the potential cytokinetic effect on the cells, as well as reducing the time for staining. However, since Celigo has one bright field and 4 fluorescent channels (Blue, Green, Red, and Far Red), fluorescence-based direct cell counting can also be easily performed.
References
- “Normalizing XF metabolic data to cellular or mitochondrial parameters”, Application Note, Seahorse Bioscience
- “Normalization of Agilent Seahorse XF Data by In-situ Cell Counting Using a BioTek Cytation 5”, Application, Seahorse Bioscience
- Riddle et al., “Expansion capacity of human muscle progenitor cells differs by age, sex, and metabolic fuel preference,” Am J Physiol Cell Physiol, doi: 10.1152/ajpcell.00135, 2018

