ELISpot is a highly sensitive, microplate-based assay for the detection and enumeration of cytokine secreting cells. It is often used to identify cytokine secretion by antigen-activated T cells and antibody secretion by B cells from peripheral blood or spleen cell preparations. T-Cell immunity assessment is a critical component of elucidating immunotherapies in cancer, infectious and auto-immune disease Zhang and Caspell et al. 2009.
What Research Areas Utilize ELISpot?
With improved ability to effectively cryopreserve peripheral blood mononuclear cells (PBMCs) Kreher, 2003, it has become possible to reliably test T cell responses in patient-specific samples.
Immuno-oncology
- Measure tumor-specific T cell responses to tumor antigens
- Measure cancer vaccine-induced antibody responses
- Profile T cell subsets
Infectious disease
- Vaccine development: measure immune responses to the vaccine
- Viral infection monitoring and treatment
ELISpot Assay Principle
- Capture antibody specific for the analyte is coated onto a PVDF-membraned microplate
- Stimulated cells are added into the wells and incubated
- Immobilized capture antibody binds secreted analyte in the immediate vicinity of secreting cells
- Cells and unbound substances are washed away
- Detection antibody and substrate are added (either colorimetric or fluorescence)
- Colored or fluorescent spots form at the sites of cytokine localization
- Each spot represents an individual analyte secreting cell
- Spots are counted either manually, or with an automated ELISpot reader
Why is Accurate Cell Counting Important for ELISpot?
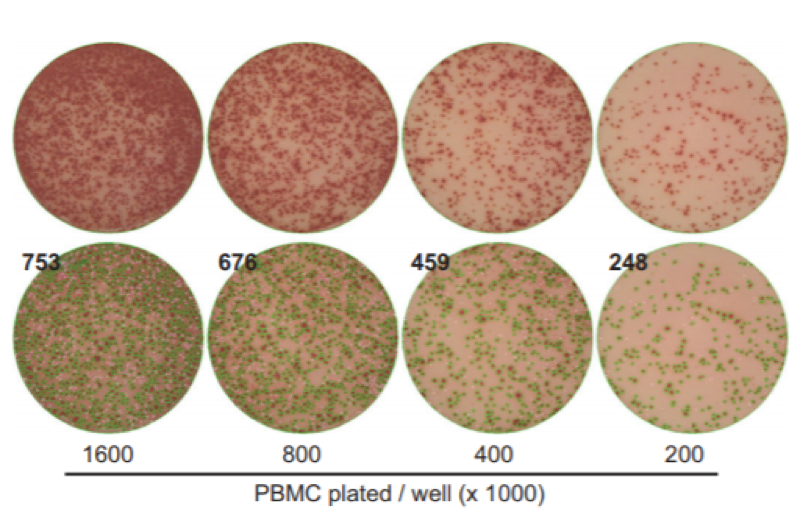
Most ELISpot experiments are performed using PBMCs, either freshly prepared or cryopreserved cells. In the particular example of T Cell activation by antigen presenting cells, a direct interaction between a T-cell and an antigen presenting cell is required to generate a “spot”, therefore an optimal density of T Cells is required to produce linear results (1 spot per bound cell). Zhang and Caspell et al. (2009) showed that in such an experiment linear results were achieved only within the range of 5×10^4 and 4×10^5 PBMCs per well.
Fig 1 Adapted from Zhang and Caspell et al., 2009. PBMC maximal density for ELISpot is shown to be 400,000 cells per well. Beyond this and spots begin to merge making it impossible to have one defined spot per bound cell.
- Number of cells added to the wells need to be optimized.
- If too many cells are added, density of spots per well will be high, which makes quantification challenging (as spots merge together becoming indistinguishable from each other).
- If too few cells are added, there will not be enough number of spots per well for reliable, statistically reproducible data.
- It is recommended to make 2-fold serial dilutions of cells to determine the optimal number of cells that will result in distinct spots.
- A high viability of sample is critical for the success of the assay
“The presence of dead cells in PBMC preparations can impact the functioning of both the T and B cell ELISPOT assay. A cut-off of ≥ 80% viability is recommended as a threshold.” https://www.ucytech.com/frequently-asked-questions-elispot
What Method Should be Used for Accurate Cell Counts of PBMCs?
Traditionally labs perform cell counts either manually or by automated cell counter using Trypan Blue to determine cell number and viability after samples have undergone RBC lysis or Ficoll separation. Two problems occur
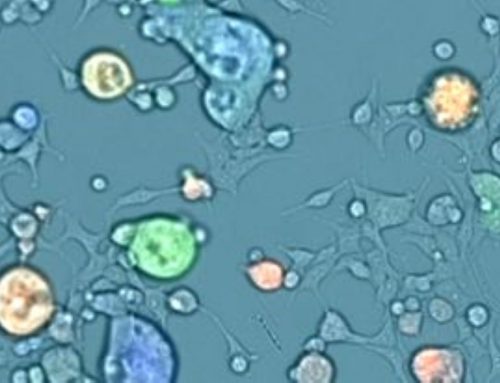
- Commonly used methods of RBC removal are often incomplete
- Trypan Blue does not distinguish nucleated (PBMCs) cells from non-nucleated cells (RBCs)
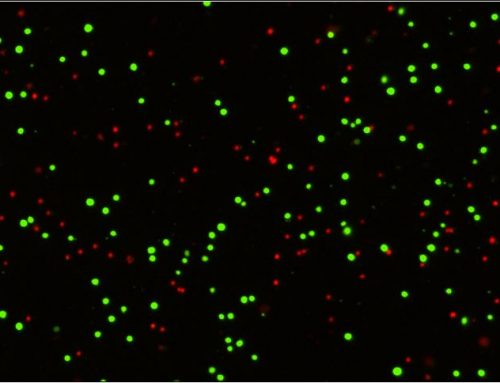
This phenomenon was detailed by Chan and Laverty et al. (2013). The most accurate method for true viability and concentration measurement of small immune cells and PBMCs is using fluorescence-based nuclear staining for live and dead nucleated cells thereby guaranteeing exclusion of RBC contamination and accurate total nucleated cell counts for the down-stream ELISpot assay.
Scott McMenemy, Nexcelom Bioscience and Dr Ning Lai, Nexcelom Bioscience
Chan and Laverty et al., J Immunol Methods. 2013 Feb 28;388(1-2):25-32.
Zhang and Caspell et al., J Immunotoxicol. 2009 Dec;6(4):227-34